Several experimental techniques are aimed at studying protein structure, which in turn can provide precious insights about molecular functions. Data are often rationalized against available high resolution structures obtained by X-ray crystallography or NMR. This can be difficult because (1) the protein conformation in an experiment might differ from its reference atomistic structure, (2) the data might only be explained by multiple conformations, and (3) the data obtained through different experiments might be ambiguous.
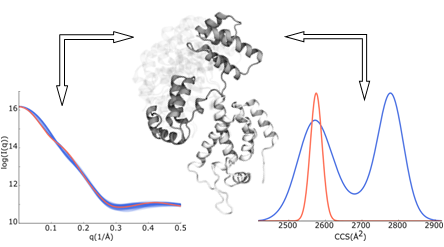
We are developing methods to compare data from ion mobility, SAXS, and chemical cross-linking with protein molecular structures, and incorporate molecular flexibility in the process.
Representative articles:
- M. Landreh, C. Sahin, J. Gault, S. Sadeghi, C.L. Drum, P. Uzdavinys, D. Drew, T.M. Allison, M.T. Degiacomi , E.G. Marklund (2020), Predicting the shapes of protein complexes through collision cross section measurements and database searches, Analytical Chemistry
- Degiacomi, M.T. , Schmidt, C. , Baldwin, A.J., Benesch, J.L.P. (2017), Accommodating Protein Dynamics in the Modeling of Chemical Crosslinks, Structure