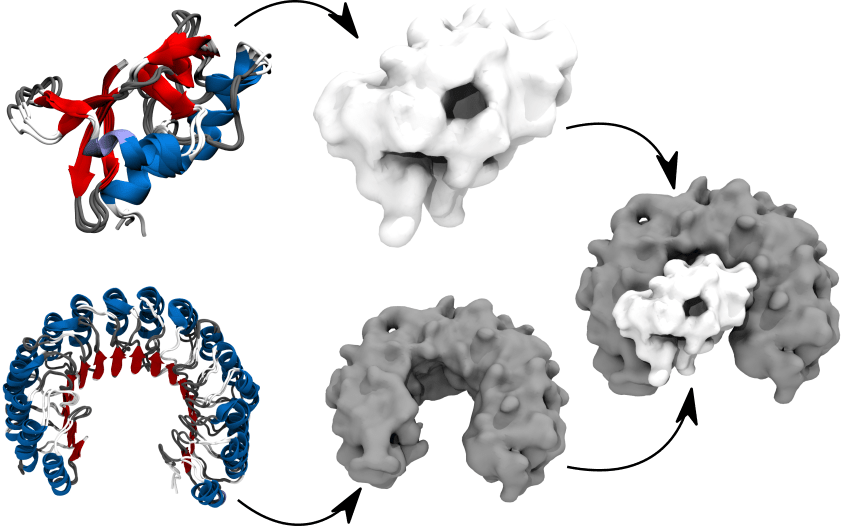
JabberDock tackles the problem of protein-protein docking while accommodating for rearrangements upon binding including side chain reorientations and backbone flexibility. To this end, JabberDock leverages Spatial and Temporal Influence Density (STID) maps, a representation of proteins surface, electrostatics and local dynamics. Proteins represented as STID maps are docked by maximising their surface complementarity using the POWer optimization engine.
DOWNLOAD
JabberDock is freely available for download on our group’s Github repository:
REFERENCE
When using JabberDock in your work, please cite the following publications.
Membrane proteins: L.S.P. Rudden & M.T. Degiacomi (2021). Transmembrane protein docking with JabberDock, Journal of Chemical Information and Modelling