Protein docking essentially involves taking the atomic structure of two proteins, generating a collection of possible molecular arrangements, and scoring their quality to determine the best one. However, the vast majority of arrangements will feature proteins that either overlap of do not touch. As these cases are uninteresting, generating them and assessing their quality is a waste of time.
Enter shape tracing. In collaboration with the Chris Willcocks group in Durham’s Computer Science Department (in particular, shout-out to Adam Leach), we combine our protein volumetric representation (STID maps) with a generalisation of ray racing capable of detecting the collision of arbitrary three-dimensional shapes.
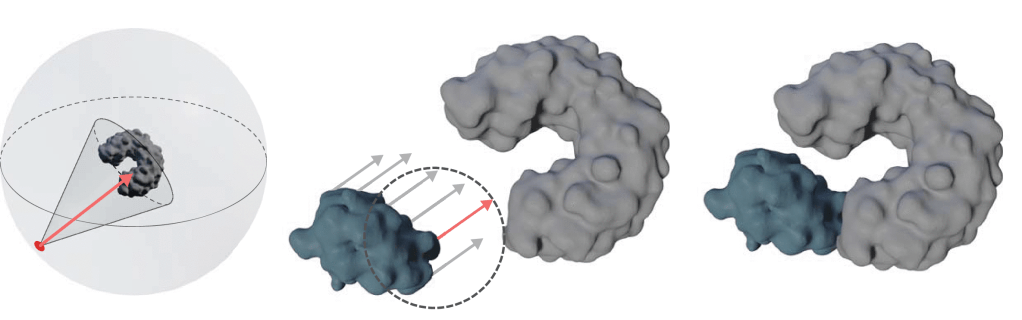
This enables us to very quickly generate arrangements of proteins that are in contact but do not overlap, speeding up the search for the most suitable model.