A fully funded studentship in the area of machine learning for computational biophysics is available in our group, starting October 2022.
All living organisms contain millions of proteins; biopolymers that fold into three-dimensional biologically active structures playing a vital role in the regulation of life and diseases. Research has seen a lot of focus on determining the atomic structure of different proteins. However, the flexible movement of these biopolymers plays a crucial role in their biological (mal)function.
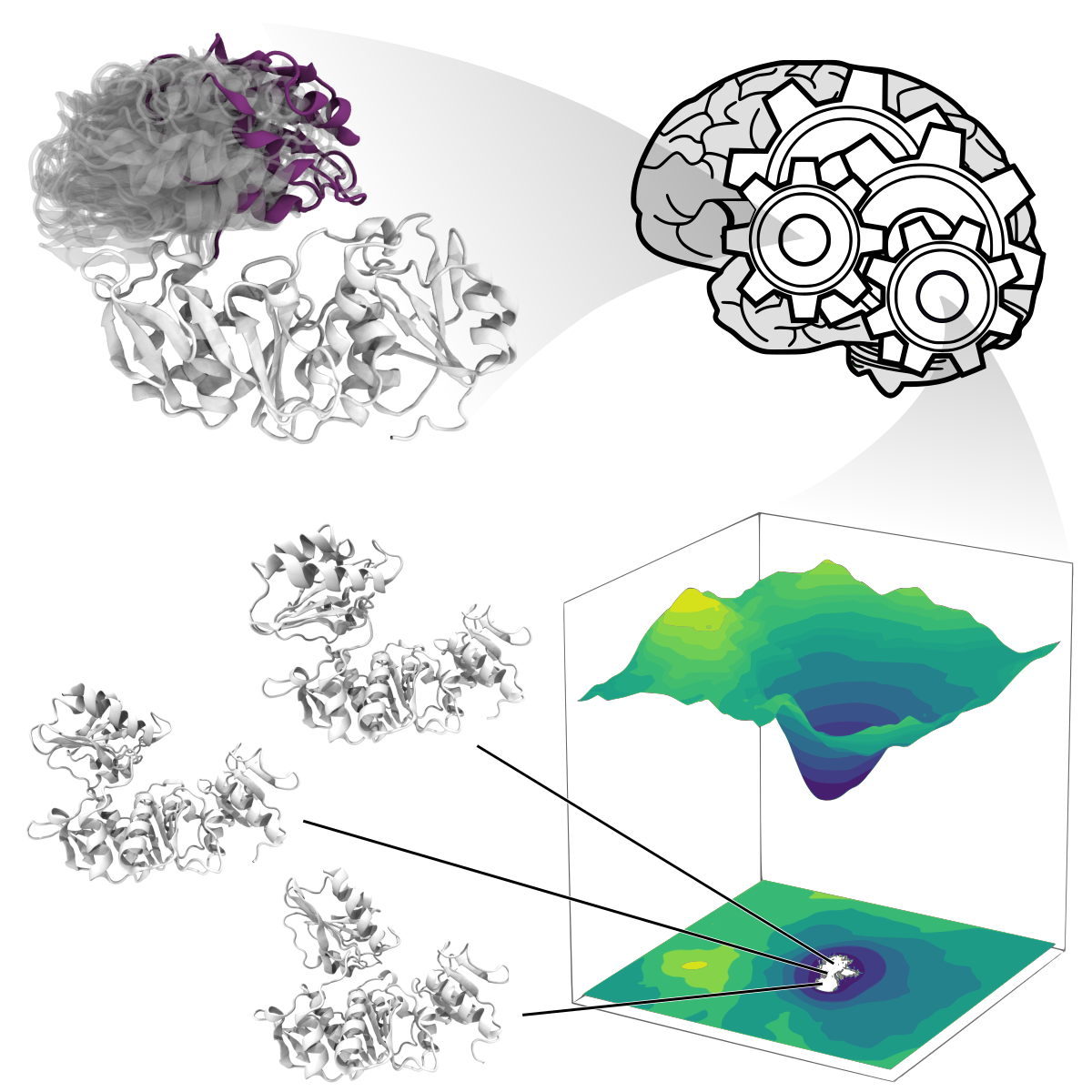
In recent years, machine learning has been revolutionizing the way we interpret data in many scientific areas. For example, the deep neural network AlphaFold2 can predict the 3-dimensional structures of proteins, whose shape is not known experimentally. In our research, we have designed a deep neural network that can also learn an ensemble of structures of specific proteins from molecular simulations. This project builds upon this breakthrough.
In collaboration with the Willcocks group (Department of Computer Science), the student will develop a general neural network capable of learning and predicting the dynamics of any protein. The neural network will be trained with existing and new data you will produce from molecular dynamics simulations. Applications of this work are vast, ranging from understanding the effect of genetic mutations in cancers to informing the design of proteins to carry out a desired function.
Do you have a background in physics, computer science, chemistry, biology, or related discipline, and are keen to develop your computational skills to address biomolecular problems? Then please direct any informal enquiries to Matteo (matteo.t.degiacomi@durham.ac.uk). More information on the application process can be found here. Applications will be considered until the 3rd April.

 In related news, in collaboration with
In related news, in collaboration with 